- Home
- Angry by Choice
- Catalogue of Organisms
- Chinleana
- Doc Madhattan
- Games with Words
- Genomics, Medicine, and Pseudoscience
- History of Geology
- Moss Plants and More
- Pleiotropy
- Plektix
- RRResearch
- Skeptic Wonder
- The Culture of Chemistry
- The Curious Wavefunction
- The Phytophactor
- The View from a Microbiologist
- Variety of Life
Field of Science
-
-
Change of address7 months ago in Variety of Life
-
Change of address7 months ago in Catalogue of Organisms
-
-
Earth Day: Pogo and our responsibility10 months ago in Doc Madhattan
-
What I Read 202410 months ago in Angry by Choice
-
I've moved to Substack. Come join me there.1 year ago in Genomics, Medicine, and Pseudoscience
-
-
-
-
Histological Evidence of Trauma in Dicynodont Tusks7 years ago in Chinleana
-
Posted: July 21, 2018 at 03:03PM7 years ago in Field Notes
-
Why doesn't all the GTA get taken up?7 years ago in RRResearch
-
-
Harnessing innate immunity to cure HIV9 years ago in Rule of 6ix
-
-
-
-
-
-
post doc job opportunity on ribosome biochemistry!11 years ago in Protein Evolution and Other Musings
-
Blogging Microbes- Communicating Microbiology to Netizens11 years ago in Memoirs of a Defective Brain
-
Re-Blog: June Was 6th Warmest Globally11 years ago in The View from a Microbiologist
-
-
-
The Lure of the Obscure? Guest Post by Frank Stahl13 years ago in Sex, Genes & Evolution
-
-
Lab Rat Moving House14 years ago in Life of a Lab Rat
-
Goodbye FoS, thanks for all the laughs14 years ago in Disease Prone
-
-
Slideshow of NASA's Stardust-NExT Mission Comet Tempel 1 Flyby15 years ago in The Large Picture Blog
-
in The Biology Files
Mystery Micrograph #21
Sunday Protist -- Rostronympha and latest parabasalian taxonomy
I can't find the original description at the moment, but vaguely recall having searched for it ambitiously once and failed miserably. It's supposed to be (Duboscq, Grassé & Rose, 1937), with a later mention in PP Grasse, A Hollande (1963) Ann. Sci. Natur. Zool. Ser "Les flagelles des genres Holomastigotoides et. Rostronympha". Might as well order it sometime...
And now onto a really nice taxonomic summary of parabsalians by Cepicka, Hampl and Kulda 2009 in Protist:
Ok, I must run...will definitely get back to parabasalians in much more detail later. Some people in our department happen to be rather obsessed with them, and obsession can be contagious. But for now, feel free to join me in salivating over that really sweet diagram!
Right, finals...
Source:
Cepicka, I., Hampl, V., & Kulda, J. (2010). Critical Taxonomic Revision of Parabasalids with Description of one new Genus and three new Species Protist DOI: 10.1016/j.protis.2009.11.005
Social onychophorans!
Onychophorans (velvet worms) are fucking adorable. Now, whether they are more or less adorable than, say, hypotrich ciliates or Apusomonas proboscidea, is open to debate (I remain loyal to my

Something about onychophoran morphology resonates quite nicely with our innate aesthetic senses...or maybe it's just me. Some of them have really pretty patterns too, or come in absolutely bizarre colours. (Mayer & Herzsch 2007 BMC Evol Biol)
A while back someone was waxing poetic about social spiders in class, which led me on quite an adventure. Since I had something very important to do that night, like an exam the following day or something, I got a lot of procrastination done: read about various social spiders (who also have an interesting story of evolutionary dead ends and conflicting levels of selection; oh, and a species with observed cooperative transport of large prey -- apparently fairly rare in arthropods), made my way to social pseudoscorpions (some of them apparently disperse by riding large insects like bugs or beetles), and then I hit upon this paper:
Social behaviour in an Australian velvet worm, Euperipatoides rowelli (Onychophora: Peripatopsidae) (Reinhard & Rowell 2005 J Zool)Social behaviour in onychophorans? Seriously!? On a second thought, why the hell not? And then came a complete overload of cute that could've only been enhanced by better images...velvet worms who cuddle!
As cuddly as they may seem, these guys also have a strict social hierarchy involving an alpha female. Reinhard and Rowell (2005) describe a feeding process where a cricket was thrown into the petri dish and attacked by the adult onychophorans (who trap their prey with sticky salivary secretions). After subduing the cricket, the first female fed on the prey for nearly an hour, biting and chasing off any other individual that would approach. After that hour, other females were allowed to feed, and then eventually males and juveniles. Most of the males were feeding after the females left. A feminist's paradise.
The interactions between individuals were observed and classified into dominant vs. subordinate: biting and chasing were done by the dominant individual (with the subordinate fleeing) whereas climbing was done by the subordinate and up to the decision of the dominant whether or not to be tolerated:
Juveniles were generally left alone and tolerated. Meanwhile, the adults were involved in a constant display of aggression and submission. Females were dominant to the males. When groups of onychophorans from different logs (thus, different social groups) were pairs, individuals of both groups acted aggressively to each other, and despite the males insisting on climbing, no aggregations were formed as they were ruthlessly rejected. Thus, the social groups are stable and at least these onychophorans seem to be capable of kin recognition.
The dominance hierarchy seemed largely size-dependent, with smaller females almost always being subservient to the larger individuals. It is believed that as in many other instances of sociality, social behaviour here aids in the cooperative capture of large prey. Curiously, the strict hierarchy when it comes to feeding, with the alpha female hoarding the entire prey, is not known in any other invertebrate.
Onychophoran behaviour doesn't receive much attention, perhaps at least partly due to the onychophoran's idea of a perfect habitat not coinciding all too well with that of an ethologist: velvet worms love cold, damp places. So it wouldn't be too surprising if an entire group of social species were eventually discovered, perhaps even with separate origins. In fact, Reinhard and Rowell (2005) state that it is not even known whether sociality may be common for onychophorans in general. On the topic of behaviour, despite their cute and almost fluffy appearance, onychophorans can also be quite vicious. This one devoured a spider bigger than itself:

Sticky spit vs. sticky silk. Quite surprisingly, the spit won this battle. (while checking whether this spider actually produces silk, found out that apprently tarantulas secrete adhesive silk from their feet...)

Onychophoran nervous system. Pseudocoloured to reflect the nerve depth in the confocal projection. Parts of the nervous system arise in a segmented fashion (eg. leg innervation), parts are repeated but not in a segmented way, and they also lack segmental ganglia as those in arthropods. Thus, onychophorans are slightly segmented in some respects, if you will, but still quite different from annelids and arthropods. This image really needs to be submitted to Nikon Small World... (Mayer & Whitington 2009 Dev Biol)
Onychophorans, conveniently branching between annelids and arthropods, seem to have little going on in the way of nervous segmentation. It has been first interpreted as having a reduced segmental nervous system with the ventral nerve cord bearing relics of reduced ganglia. The ring nerve bundles are absent in the leg regions (and present between them) in a segmentally-appearing fashion; however, Mayer & Whitington (2009) have shown that the number of bundles between each leg pair varies and that unlike in arthropods, ventral organs do not coordinate neural development, as they develop fairly late relative to the nervous system. Curiously, the function of ventral organs in onychophoran development seems to be poorly understood.
Thus, segmental ganglia seem to have evolved in annelids and arthropods separately, adding further support to the Ecdysozoa idea (a debate I know next to nothing about...). The story of protist phylogeny has shown that convergence is much more rampant than we'd like to think, so it would be quite interesting if the nervous system of annelids and arthropods is also a case of convergent evolution, further showing how outright dangerous it is to rely on morphology for phylogenetic reconstruction. Metazoan phylogeny is a mess. While trying to get a grip on invert phylogeny for a class, I found it rather daunting how much I'd have to learn in order to make any sort of personal judgement on the matter. While protists are enough to keep one busy and fascinated (and overwhelmed) for many lifetimes, it is still fun to occasionally wander outside one's field and check out the daunting questions others have to deal with.
Besides, prior to taking a course in developmental biology, I actually had quite an interest in it, before it was quelled mercilessly by having to memorise structures of the 72h chick embryo. Without any evolutionary context. I think studying unicellular development would be useful for those studying multicellular development, as there must surely be ultimate convergence between many processes and structures. After all, there's only so many ways one can establish polarity, regulate morphology, grow, reproduce, etc. Comparative study of the developmental biology of the various eukaryotic (and prokaryotic!) supergroups and phyla therein would surely be a fun field once enough is known about organisms that are NOT mice, mustard weed, baker's yeast and fruit flies.
*OMG there are also eusocial thrips. With soldiers.
** I wonder if the traditional zoologists' repulsion towards anything molecular may be part of it...
References
Dias, S., & Lo-Man-Hung, N. (2009). First record of an onychophoran (Onychophora, Peripatidae) feeding on a theraphosid spider (Araneae, Theraphosidae) Journal of Arachnology, 37 (1), 116-117 DOI: 10.1636/ST08-20.1
Mayer, G., & Harzsch, S. (2007). Immunolocalization of serotonin in Onychophora argues against segmental ganglia being an ancestral feature of arthropods BMC Evolutionary Biology, 7 (1) DOI: 10.1186/1471-2148-7-118
Mayer, G., & Whitington, P. (2009). Neural development in Onychophora (velvet worms) suggests a step-wise evolution of segmentation in the nervous system of Panarthropoda Developmental Biology, 335 (1), 263-275 DOI: 10.1016/j.ydbio.2009.08.011
Reinhard, J., & Rowell, D. (2005). Social behaviour in an Australian velvet worm, Euperipatoides rowelli (Onychophora: Peripatopsidae) Journal of Zoology, 267 (01) DOI: 10.1017/S0952836905007090
What songbirds can teach us about language: Does syntactic complexity require selection?
Here's another story of how alleviated selective pressures can enable increased complexity, without said increase in complexity needing to be driven by positive selection; the paper later relates this to its implications for language evolution:
Ritchie & Kirby 2005 Evol Ling Comm Selection, domestication, and the emergence of learned communication systems
(Also see Ritchie & Kirby 2007 Emergence of Communication and Language)
In a prior study, the Bengalese finch was found to have a more complex song syntax than its wild ancestor (the white-backed munia); furthermore, while the finches could learn the songs of munia, the latter could not effectively learn the complex songs of the finches, suggesting that a part of the capability was physiological. The author of that prior study, Okanoya (2002), argued that the song complexity in the Bengalese finch was driven through sex selection, as the more basic pressures (food and predation) were relieved by domestication, enabling sex selection to finally drive up the complexity; furthermore, this would have been an honest signal of the male’s fitness, as a fitter bird could produce a more complicated song.
A competing hypothesis by Deacon agrees that song complexity is kept low in the wild munia through selective pressure, but claims that the lifting of basic selective pressures after domestication enabled the songs to get more complex through other means, namely drift. Thus, processes that previously paled in comparison with the selective pressures against excess song complexity became prominent, such as the effect of songs heard at an early age and mnemonic biases; that is, songs with a more regularised syntax may be easier to recall. Deacon further extends this concept to the evolution of human language; he calls the concept “selective masking”. In short, complexity may arise in the finches’ songs without being driven by direct selective pressure.
Ritchie and Kirby set out to test the competing hypotheses through computational modeling; long story short, a bunch of learning filters are set up amid evolutionary models, and the simulation is run trough three phases: 1) Population is filtered to have a particular song type; variation is reduced (modeling the case among wild munia. 2) The population, having learned (and become “attached” to) a particular kind of song, is now bombarded with a bunch of random songs, and demonstrates resistance to be affected much by it: the simulated birds still learn the ‘correct’ song over incorrect ones. 3) Selective pressure is alleviated altogether by simply ceasing to calculate the fitness values. This simulates domestication. Population was once again bombarded with random songs.
Complexity was initially defined by Okanoya as the number of unique song notes divided by the number of unique note-to-note transitions (aka ‘Song Linearity’); Okanoya found this ratio to be lower in the Bengalese finches than the wild munia, meaning their songs were more complex (less ‘linear’). Ritchie & Kirby’s simulation also yielded similar results; though they argue that a completely random song would have the maximum complexity by such measures. Additionally, they also used Grammar Encoding Length, or the number of bits required to describe a [in this case, Probabilistic] Finite State Machine, which was used to model song learning. [Now the structural linguistics and information theory loses me completely...]. Turns out, in phase 3, the grammar encoding length did increase and the linearity did go down, supporting the increase in song complexity after domestication.
Most importantly, their simulation showed that song complexity can increase in the absense of direct selective pressure, as selection was eliminated altogether in phase 3. This suggests that [once again,] one need not necessarily evoke often-absurdly-complex selection stories (like sex selection and ‘honest signals’) to explain the song complexification in Bengalese finches. Furthermore, these results can be extrapolated further to linguistic evolution, suggesting that perhaps not all of syntax complexification requires selective pressures behind it. In fact, the eliviation of such pressures can allow more complex syntax to arise. As a sidenote, it has been observed that the rise of writing resulted in higher complexity of clause embedding, and this complexity also rose gradually, not immediately after writing systems first appeared (Karlsson in Sampson et al. 2009 Language Complexity as an Evolving Variable). This can also be seen as a case of the lifting (aka ‘masking’) of a selective constraint resulting in allevated complexity, in this case probably not particularly adaptive either. One can convey complex ideas just as well, and in some cases better, without the [ab]use of intricate clausal embedding…
Ritchie and Kirby conclude with an idea that perhaps one mustn’t look for selective advantages of elaborate syntax found in human language, but instead investigate what may have prevented syntactic elaboration from arising in the past – what selective pressure may have been alleviated, and what may have caused them?
Ritchie GRS, & Kirby S (2007). A Possible Role for Selective Masking
in the Evolution of Complex, Learned
Communication Systems Emergence of Communication and Language, 387-401 DOI: 10.1007/978-1-84628-779-4_20
(Ritchie + Kirby 2005 turns out to be a draft for the 2007 paper, and I'm far too sleepy to reference that properly as it's too complicated...)
Mystery Micrograph #20
You guys are too good at this...next!
(Oh, and I am working on writing up the past ones... just so you know -- they're coming, eventually...)

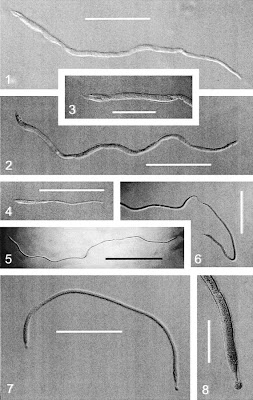
Scalebar = 500um; to be referenced later
Sunday Protist -- Gymnodinium catenatum: Like beads on a string
Moss Microforay Part 1: Assorted critters
Moss was sampled on a coastal mountain on the San Francisco peninsula. As the sampler is a labrat and therefore not particularly sample-savvy, said specimen was abandoned for three weeks only to be rehydrated a couple days prior to imaging. Thus, there is a high probability that much of the really cool stuff died off before I got around to it. Conversely, some of that cool stuff may be damn good at encysting, and is now perhaps freshly-excysted.
Saving the Euglyphids for the next installment. Mwahaha. Let's go over an assortment of random non-Euglyphid organisms first. It would be nice to get some help with ID from the experts lurking here *wink*
Ok, first off -- flagellates. That is, glimpses of the few flagellates that weren't moving so ridiculously fast they were but a blur on the picture. There seemed to be a lot of bicoesid-like loricate things (or vaguely look like them anyway):

That cell may have been damaged somehow, or is it actually being amoeboid-ish? Can bicoesids do that? My understanding is they tend to be rather rigid, but I'm not aware of any other loricate flagellate besides Dinobryon and choanoflagellates. This thing is definitely neither of those.
This specimen appeared attached along the length of the flagellum at the top of the picture.

This one looks more like a bicoesid. As do the next three:



This one too, probably, though the lorica can't be seen. There are aloricate bicoesids as well.

Bicoesids are actually borderline suitable for microphotography with a slow camera due to their habit of attaching to things. Seriously, sessile organisms are really nice that way...
More potentially cool flagellates that were moving around too much to get a decent shot:



This was a sequence following a strange amoeboflagellate-looking thing(s) -- any idea what this may be? A cercomonad of sorts?

Same cell(s) in all of the images. Was shot in a short timeframe (maybe a minute or two), so highly doubtful that cell divison could've occured within that time. Or maybe not?
Amoebozoan time! May ever try ID'ing some...before doing that, I feel compelled to draw your attention to Smirnov & Goodkov 1999 Protistology, a wonderful overview of and reference for amoeboid morphotype diversity -- amoebae do have shape! Another good source for ID'ing amoebae is Smirnov & Brown 2004 Protistology, and an associated site (though sadly not maintained since 2003ish =( ); I will be relying on those to see how far I can go:

This one seems to be flabellate with adhesive uroidal filaments; may be Flabellula (something off about that), Stachyamoeba (perhaps? esp. this picture); Rosculus seems to lack prominent uroidal filaments as well as seemingly a Heterolobosean from what I've gathered. Could also be of the fan-shaped morphotype but those don't seem to leave trailing adhesive filaments that much. I'll go with Stachyamoeba for now, hopefully no one finds that offensive...
Ok, the next one is probably polytactic or dactylopodial by morphotype.

Candidates: Amoeba (too small), Chaos (doesn't seem multinucleate and huge), Deuteramoeba, Thecochaos (not wrinkly enough), Polychaos (something meh about that too). As for dactylopodial candidates, Paramoeba is looking better, as is Korotnevella. As the former is much more commonly mentioned, I'll go with Paramoeba for this one.
By the way, the prominent round thing in the middle is most likely the contractile vacuole; note how it obvious changes size during optical sectioning, especially in the first specimen. Also, amoebae aren't always amoeboid -- many species take upon 'floating forms' which tend to look like spiky balls. Some amoebae are apparently quite at ease with being planktonic too, like this vicious rotifer-munching Difflugia mentioned earlier.
Amoebae can actually move pretty quickly -- this one is fuzzy due to motion blur.

Morphotype: Lens-like (Smirnov & Brown 2004); candidate: Cochliopodium (and here). Really does look like Cochliopodium; perhaps I got at least one right!
A few more fun amoebae:



In the last one, note how the clear pseudopodia emphasising the contrast between ecto- and endoplasm -- the latter being fairly gelatinous and containing the 'stuff' of the cell; the former, full of actin, responsible for cell motility.
Arcellid amoeba: (proteinaceous test -- note texture)

Odds and ends: First off, ???:


What I think might be a developing nematode egg case (elsewhere there were very similar things with nematodes inside):

Some fungal thing? (germinating spore?)

And last but not least for today, a bacterial filament:

Ok, back to cramming...stay away, internet! Must know everything there is to know about invert biol by Monday...aaaah~! Also, apparently metazoan phylogeny is in about as much of a mess as the the Tree of Eukaryotes. Kind of creepy...
In defense of constructive neutral evolution - Part II
Part I
Part II
-Neutral evolution is relevant
-Evolution lacks foresight; it can neither anticipate nor respond
-Clarifying some terminology: two types of function, positive vs. negative selection
-Rise of complexity through non-adaptive means
-An example of constructive neutral evolution at work: loss of group I intron self-splicing
Part III
-Further examples of constructive neutral evolution
-Discussion of what sparked this argument: Evolution of ciliate nuclear dimorphism
Neutral evolution is relevant
Again, apologies for stating the obvious, but apparently even some prominent
"The key point that blunts the Gould and Lewontin critique of adaptationism is that natural selection is the only scientific explanation of adaptive complexity." [p.6; emphasis mine]First off, it's kind of cute that a couple psychologists seem to think they can so easily outright dismiss a point made by evolutionary biologists. I mean, seriously, I find it adorable. On that note, I'm now gonna write a book chapter debunking generative linguistics, because, well, my buddy says they're wrong. I wonder how much Pinker's evolutionary views have been shaped by Dawkins et al. Their camp is rather influential outside evolutionary biology, and while many claim that panadaptationism is a strawman -- and even in biology that point is debatable -- outside evolutionary biology, panadaptationism is alive and well. Part of the reason is that adaptationist stories are written in popular books, while pluralistic approaches largely remain hidden in the likes of Molecular Biology & Evolution, Biology Direct and Journal of Molecular Evolution. Researchers working in applied evolutionary fields have likely never heard of them.
Back to Pinker & Bloom, first off it's quite a tautology to claim that adaptive complexity evolves through adaptation. Well, yes, you JUST labelled it adaptive. In the immortal words of 4chan, long cat is loooooong. Casting that aside, they've falled for a false dichotomy. You would think someone as smart as Steven Pinker and Paul Bloom wouldn't fall for it. But they did. Why does it have to be one or the other? Why adaptation or spandrel? Especially when we speak of highly complex systems -- is it not in the definition of complexity that they consist of multiple components? How likely is it that all of them are adaptive or neutral or maladaptive? Is it even remotely productive to reduce a system so drastically as to label it simply as an adaptation? What does that even mean, besides stating the obvious? 'Adaptation' has got to be one of the more utterly useless terms in evolution biology, at least the way it's abused today.
Letting Pinker & Bloom speak further:
"Adaptive complexity" describes any system composed of many interacting parts where the details of the parts' structure and arrangement suggest design to fulfill some function. The vertebrate eye is the classic example. The eye has a transparent refracting outer cover, a variable-focus lens, a light-sensitive layer of neural tissue lying at the focal plane of the lens, a diaphragm whose diameter changes with illumination level, muscles that move it in precise conjunction and convergence with those of the other eye, and elaborate neural circuits that respond to patterns defining edges, colors, motion, and stereoscopic disparity. It is impossible to make sense of the structure of the eye without noting that it appears as if it was designed for the purpose of seeing -- if for no other reason that the man-made tool for image formation, the camera, displays an uncanny resemblance to the eye. Before Darwin, theologians, notably William Paley, pointed to its exquisite design as evidence for the existence of a divine designer. Darwin showed how such "organs of extreme perfection and complication" could arise from the purely physical process of natural selection." [p.6 cont'd]Holy fucking crap, argument from design, for selectionism! Impressive. Yes, you guys just totally pwned Gould with the vertebrate eye. He was unaware of its very existence. Eyes don't preserve well in the fossil record, you see? Eyes look designed, therefore selection. Great. I'll let them finish...
"The essential point is that no physical process other than natural selection can explain the evolution of an organ like the eye." [p.6]AFAIK, Gould never denied selection!!! He was a brilliant biologist who thoroughly understood evolution, unlike some recent drama queens. What Gould argues is that a) not everything is an adaptation and b) adaptation is neither the sole nor the most important 'force' in evolution (nor is it actually a 'force' of any sort...). Furthermore, while I am unaware of Gould's opinions about The Eye, and am currently too lazy to research the topic, to me it seems highly implausible that the vertebrate eye evolved solely through selection. In fact, considering that selection is a purifying, not driving, 'force' -- that is, selection simply removes the not sufficiently fit -- it is curious to see where Pinker & Bloom think the 'material' for the selective evolution of the eyes comes from. Surely they're not insane enough to believe that the thing evolved entirely through point mutations each making the eye progressively slightly better and better and suddenly, veeeery gradually, ta-da: The Eye!
On that note, I wonder if there's a strong link between selectionism and gradualism. In a sense, one does kind of have to believe the above point-by-point scenario to explain how anything arises purely by selection. To anyone with the slightest inkling of how genes and genomes work, such a view is obviously absurd. Although considering how Dawkins has already enlightened us that molecular biologists may or may not be -reputable- biologists...
I'm still amused by how remarkably cute it is of Pinker & Bloom to know so much about the details of evolution that they can make bold statements like the last sentence cited above. That's one strong statement!
tl;dr Some rather reputable and smart people still fall for the false dichotomy where something is either purely adaptive or purely 'random'. Selection is not the sole source of order (Lynch 2007 PNAS; you might as well head over and read that paper by now)
So if selection is not the only source of order, what else could be? How can neutral forces (coupled with negative selection) result in an increased complexity? Ah, nearly time for constructive neutral evolution. But prior to that, one more thing to take care of...
Evolution lacks foresight; evolution is not engineering
First of all, one must note that a modern function of any given system may not necessarily have been there at the initial stages of evolution. In fact, it is often counterproductive to even assume so. A nice example would be diatom sex.
Upon each division, one of the diatoms gets smaller. The cell is surrounded by two valves: a larger top valve and a smaller bottom one. During division, the bottom one actually becomes the top in the daughter cell. After several generations, some diatom offspring get a little on the small side. Luckily, sex makes them bigger; ie. they produce gametes (or swap nuclei), fuse and make a brand new maximum-sized cell. Jennifer Frazer wrote an awesome post on this topic.
One may look at diatoms and say: What is the function of sex? If you remove sex, the diatoms can't get bigger again, and die a miniscule death. Thus, sex must be there to get bigger!
However, one must imagine evolution as a blind step-by-step process, devoid of any foresight or momentum whatsoever. A common fallacy would be to think of the diatoms first having a problem, and then having to come up with an emergency solution to it, eg. evolve sex. This may seem obviously ridiculous in the case of something as complex as sex, but is often assumed for simpler cases, like evolving a resistance to something. Perhaps we get this picture from the antibiotic resistance examples in bacteria; however, antibiotic resistance is generally much too simple, and bacteria much too numerous, to apply this mode of evolution to everything else. It is additionally pretty rare, but we don't notice the vastly overwhelming multitude of unsuccessfulcases.
Back to our diatoms, the more plausible scenario is actually that sex was there first, and was able to correct for the problems caused by this rather suboptimal mode of mitosis. Sex enabled this peculiar life style; it did not evolve in response to it. 'Enable' is a word that must be used much more frequently in the literature. Certain situations enable certain mutations or other changes to persist or excalate into fixation (either by drift or selection). They do not actually push them there.
Again, evolution cannot anticipate.
Two meanings of 'function'
As a brief aside, it is important to note two distinct meanings of the term 'function', often used rather carelessly:
1. Selected function - what a certain trait would have been selected for in its evolutionary past.
2. Current function - based upon what would happen were the trait removed in the present.
Going back to our diatom example above, the current function of sex in diatoms can be argued as a way to increase the population size back to original levels; for inhibiting sex would remove a way for the organisms to get bigger again. However, this is not the selected function of sex, since it must have already existed before the size problem happened. (the selected function of sex, if any, remains a fuzzy, murky mess)
Positive vs. negative selection
One more brief thing to clean up: 'types' of selection
Negative (aka purifying) selection - selection acting against a trait
Positive selection - selection promoting a trait in a population, usually through competition (the type most commonly spoken of in popular media, despite being far less common to the point of relative rarity...)
Ultimately, positive selection is a form of negative selection where the appearance of a fitter form of a trait renders the rest of the variants relatively less fit, resulting in them being selected against (negative selection). Thus, selection isn't actually ever for anything, strictly speaking. Selection is a constraint, weeding out variants that are insufficiently fit. Furthermore, selection is always lurking about in background, albeit in varying intensities depending on the conditions (see Part I). That said, one must not ever conflate selection with adaptation -- they are not the same thing by any means!
Rise of complexity through non-adaptive means
I won't go into much detail here as I'm sure Arlin Stoltzfus himself will explain it much better in his upcoming posts on Sandwalk. Also, unlike me, he's actually qualified to write about this stuff. I will give a crude overview (to the best of my understanding) along with a few concrete examples. For an additional source on this, I blogged Ford Doolittle's seminar talk on the subject here. The non-adaptive evolution of complexity requires tinkering, relaxed selection (eg. due to smaller effective population size) and ratchets. Tinkering is basic (as mentioned above, evolution lacks foresight, many adaptations are far from optimal as relics of past functions are retained, etc) so I won't discuss it. Small population sizes were discussed in Part I, so now we need the final piece: Ratchets. This is where Constructive Neutral Evolution enters.
In a nutshell, chance interactions may happen to compensate for otherwise-deleterious mutations, thereby enabling them to eventually occur, thereby ultimately resulting in an irreversible dependency upon the interaction. That way, complexity (loosely defined as number of components and interactions within a specified system; as in Lukes et al 2009 PNAS) can increase without ever actually being adaptive. This mechanism stresses that not all complexity is actually 'better', and perhaps much of it may well be a case of mere bloating, much like the bureaucracy of an institution with time. For a system that is required to be constitutively functional, it is in fact easier to bloat complexity than to trim it down, partly just due to basic combinatorics: there are many more ways of being complicated than being simple, and just statistically there'd be more viable states of higher complexity.

Overview of Constructive Neutral Evolution. Modified and enhanced from older diagram here; Ford Doolittle uses something similar in his talk, so it shouldn't be too far off.
Cyt-18 and the N.crassa mitochondrial self-splicing intron
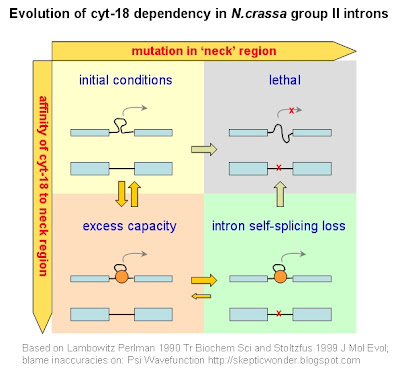
I will discuss the not-so-self-splicing intron examples in further detail in part III, but just to provide a quite example to the above point, let's look at a Neurospora mitochondrial self-splicing intron that developed a dependency on another protein. This example is based on an old-ish paper, Akins & Lambowitz 1987 Cell. It was found that mutants of cyt-18 exhibited defects in the splicing of group I (self-splicing) introns. Curiously, cyt-18 turned out to be homologous to tRNA synthetases. Further work confirmed that cyt-18 is necessary for splicing in N.crassa, and phylogenies showed it to be a derived trait. That is, in closely related lineages, cyt-18 is NOT necessary for group I intron splicing.
Now let's say this cyt-18 protein happens to bind to the neck region, and stabilise it slightly. In the grand scheme of things, the intron can splice out properly regardless of whether this protein is there, so it doesn't increase fitness in any way. However, what this protein does to is create an excess capacity in the system by enabling certain otherwise-deleterious mutations to arise in the neck region by compensating their destabilisation effects. Once such a mutation occurs (and it's bound to happen eventually), the lineage now depends upon cyt-18 for proper splicing to occur. Since the chance of the mutation being reverse is quite small, and often even less than the chance of another otherwise-deleterious mutation further fixing the protein dependency, the system is now stuck depending on cyt-18, with no adaptive advantage whatsoever. In a diagram:
References
Akins, R., & Lambowitz, A. (1987). A protein required for splicing group I introns in Neurospora mitochondria is mitochondrial tyrosyl-tRNA synthetase or a derivative thereof Cell, 50 (3), 331-345 DOI: 10.1016/0092-8674(87)90488-0